NEWS:
2016-05-15: New Release - RCaller 3.0
2015-03-13: New Release - RCaller 2.5
2014-07-15: New Release - RCaller 2.4
2014-06-07: New Research Paper on RCaller
2014-05-15: New Release - RCaller 2.3
2014-04-15: New Release - RCaller 2.2
I have received too many e-mails since i had first submitted the early versions of the RCaller. Some users found it usable so i was planning to develop a newer and enhanced version of this library. Now, i think, it is ready for testing. The 2.0.0 version of the RCaller is downloadable from
http://code.google.com/p/rcaller/downloads/list
with both compiled jar file and the source file with the directory structure of NetBeans 7.
The use of RCaller is changed after version 1.0 but it is still easy to implement, it does not need extra libraries, it is platform independent and compatible with the recent R versions.
Some new features in version 2.0.0 are:
1) Support for plots
2) Easier code generation
3) Enhanced interaction with R
Before anything, install the R package "Runiversal" by typing
install.packages ( "Runiversal" )
in R interactive interpreter. If installation is success, you are ready for calling R from Java.
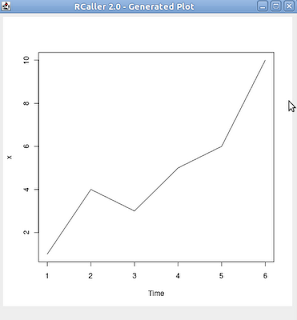
Let me explain them with some examples. Suppose that we have a double array with values of {1,4,3,5,6,10} and we want to show a time series plot with this. Firstly we import the needed libraries:
import java.io.File; import rcaller.RCaller;
We are declaring the double array:
double[] numbers = new double[] {1,4,3,5,6,10};
Creating an object of Rcaller class:
RCaller caller = new RCaller();
RCaller needs the Rscript executable file (Rscript.exe in windows) which is shipped with the R. You must tell the full path of this file in RCaller like this:
caller.setRscriptExecutable("/usr/bin/Rscript");
This is the location of my Rscript file in my Ubuntu Linux. We didn't do much thing, but this code initializes the whole thing:
caller.cleanRCode();
the cleanRCode() function of RCaller class cleans the code buffer and puts some code in it. You can browse the source code if you want to know more about the initialization. Now, we can add our double array to our R code:
caller.addDoubleArray("x", numbers);
Now we have 'x' with the value of numbers[]. Now we are creating the time series plot:
File file = caller.startPlot();
caller.addRCode("plot.ts(x)");
caller.endPlot();
Finally we are sending the whole code to R interpreter:
caller.runOnly();
With this code, Rscript runs our code but it does not return anything. After all, if we have'nt got any errors, we can handle the generated image using
ImageIcon ii=caller.getPlot(file);
or we can show it directly using
caller.showPlot(file);
The generated plot is shown below:
The source code of entire example is given below:
package test;
import java.io.File;
import javax.swing.ImageIcon;
import rcaller.RCaller;
public class Test1 {
public static void main(String[] args) {
new Test1();
}
/*
* Test for simple plots
*/
public Test1() {
try {
RCaller caller = new RCaller();
caller.setRscriptExecutable("/usr/bin/Rscript");
caller.cleanRCode();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
caller.addDoubleArray("x", numbers);
File file = caller.startPlot();
caller.addRCode("plot.ts(x)");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
} catch (Exception e) {
System.out.println(e.toString());
}
}
}
This example shows how to send vectors to R, call R function from Java and handle the result from Java. We are generating the x and the y vectors in Java and sending an "ordinary least squares" command to R. After running process, we handle the calculated residuals, fitted values and residuals from Java. I hope the example is clear enough to understand.
package test;
import rcaller.RCaller;
public class Test2 {
public static void main(String[] args){
new Test2();
}
public Test2(){
try{
//Creating an instance of class RCaller
RCaller caller = new RCaller();
//Important. Where is the Rscript?
//This is Rscript.exe in windows
caller.setRscriptExecutable("/usr/bin/Rscript");
//Generating x and y vectors
double[] x = new double[]{1,2,3,4,5,6,7,8,9,10};
double[] y = new double[]{2,4,6,8,10,12,14,16,18,30};
//Generating R code
//addDoubleArray() method converts Java arrays to R vectors
caller.addDoubleArray("x", x);
caller.addDoubleArray("y", y);
//ols<-lm(y~x) is totally R Code
//but we send the x and the y vectors from Java
caller.addRCode("ols<-lm(y~x)");
//We are running the R code but
//we want code to send some result to us (java)
//We want to handle the ols object generated in R side
//
caller.runAndReturnResult("ols");
//We are printing the content of ols
System.out.println("Available results from lm() object:");
System.out.println(caller.getParser().getNames());
//Parsing some objects of lm()
//Residuals, coefficients and fitted.values are some elements of lm()
//object returned by the R. We parsing those elements to use in Java
double[] residuals = caller.getParser().getAsDoubleArray("residuals");
double[] coefficients = caller.getParser().getAsDoubleArray("coefficients");
double[] fitteds = caller.getParser().getAsDoubleArray("fitted_values");
//Printing results
System.out.println("Coefficients:");
for (int i=0;i< span="">
System.out.println("Beta "+i+" = "+coefficients[i]);
}
System.out.println("Residuals:");
for (int i=0;i< span="">
System.out.println(i+" = "+residuals[i]);
}
}catch (Exception e){
System.out.println(e.toString());
}
}
}
I hope it works for you.

Thank you for your update for the new Rcaller.
ReplyDeleteI am using windows 7 with R 2.13 and netbeans 7.
It shows new instance ignored, assign return value to new variable.
I've been modified to Test1 test1 = new Test1();
but the result remain the same.
it just show a new window, but no plot in it.
It also shows variable ii is not used.
Will you mind to take a look at it? Did I put the right path for calling Rscript.exe?
Thank you.
Thank you
public class Test1 {
/**
* @param args the command line arguments
*/
public static void main(String[] args) {
// TODO code application logic here
new Test1();
}
public Test1() {
try {
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:\\Program Files\\R\\R-2.13.0\\bin\\Rscript.exe");
caller.cleanRCode();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
caller.addDoubleArray("x", numbers);
File file = caller.startPlot();
caller.addRCode("plot.ts(x)");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
} catch (Exception e) {
System.out.println(e.toString());
}
}
}
After I tried to use it in the Mac, it works...
ReplyDeleteI think there is some problem with my widows setup
It will be great when you have free time and you may teach us how to use it in windows. Thank you! rCaller is so useful!
we are using RCaller in our educational project. Thanks for update him.
ReplyDeletethe licence of RCaller 2.0 is LGPL by now.
ReplyDeletehttp://stdioe.blogspot.com/2011/07/about-licence-of-rcaller.html
how do i download the jar file? :)
ReplyDeletei go to the google code page but i cant seem to download it anywhere
The jar file is under the folder ../../dist:
ReplyDeleteuse this link:
http://code.google.com/p/rcaller/source/browse/#hg%2FRCaller%2Fdist
caller.addDoubleArray("x", x);
ReplyDeletecaller.addDoubleArray("y", y);
caller.addRCode("ols<-lm(y~x)");
caller.runAndReturnResult("ols");
It gives me an error for the last line saying : "Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException: Premature end of file"
I kinda use jsp pages over core java since that is what we use here...
as you mailed me, it was resolved after restarting the tomcat.
ReplyDeleteCan you please share the problem with its solutions here?
I had mailed you about some other problem which was with regards to getting the new jar file working itself. That was basically a hot deploy not working for tomcat which was solved by restarting tomcat :)
ReplyDeleteI cant seem to solve the above problem though...
So i tried setting up the entire thing in Eclipse but I still get the premature end of file result. Am I doing anything wrong with the new build? The old build used to work perfectly fine except when I had to load a new package for some functions.....
ReplyDeleteit seems there are several problems in Windows machines, i will debug the test suite in a Windows machine as soon as possible.
ReplyDeleteI tried it on linux as well with the same code as Test3 just now but I seem to get the same error :
ReplyDelete"[Fatal Error] Routput8379599187707067589:1:1: Premature end of file.
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException; systemId: file:/tmp/Routput8379599187707067589; lineNumber: 1; columnNumber: 1; Premature end of file."
I just tried it again in Linux without using an external package, working fine now....but when I try to use an external package it gives me the premature end of file exception. I guess the library ("") command throws it off a little?
ReplyDeletefirstly, for the "currupted jar file" error try to download the jar file from the link
ReplyDeletehttp://code.google.com/p/rcaller/downloads/list
this is out of the source code and uploaded manually but it may be out of date. For a fresh download, it is best to use mercurial clients for downloading source code and compile manually.
Sataract: can you please send your code which throws the parse exception? so i can solve the problem which is because of the "library()" function.
The package name is Rglpk. If i print out what I add to RCode and try it in R, it works fine...
ReplyDeletepackage RCaller;
import java.util.Random;
import rcaller.RCaller;
public class Test3 {
public static void main(String[] args){
new Test3();
}
public Test3(){
try{
Random random = new Random();
RCaller caller = new RCaller();
caller.cleanRCode();
caller.setRscriptExecutable("/usr/bin/Rscript");
caller.addRCode("library(Rglpk)");
caller.runOnly();
caller.addRCode("obj <- c(2,4,3)");
caller.addRCode("mat <- matrix(c(3,2,1,4,1,3,2,1,2),nrow = 3)");
caller.addRCode("dir <- c(\"<=\",\"<=\",\"<=\")");
caller.addRCode("rhs <- c(60,40,80)");
caller.addRCode("max <- TRUE");
caller.addRCode("mylp <- Rglpk_solve_LP(obj, mat, dir, rhs, max = max)");
caller.addRCode("myall<-list(optimum=mylp$optimum,solution = mylp$solution)");
caller.runAndReturnResult("myall");
}catch(Exception e)
{
e.printStackTrace();
//System.out.println(e.toString());
}
}
}
Sataract:
ReplyDeletethe line "caller.runOnly();" is unnecessary. Try running your code without this line.
hey it worked perfectly just now. Thanks a lot again.
ReplyDeletei know it is still problematic in Windows systems, i will correct those problems as soon as possible.
ReplyDeleteAs users said, it works perfect on Linux and Mac systems.
I have just submitted a new Jar file that seems okay on windows systems. please check it.
ReplyDeleteHi! I just stumbled upon your very useful program RCaller. I installed it and ran the test files without any problem. However, for some unknown reason, it does not execute my RSript. Here is my Java code:
ReplyDeleteRCaller r = new RCaller();
r.cleanRCode();
r.setRscriptExecutable("…");
r.addRCode("source(\"… (some path to RScript file)");
r.runOnly();
The RScript file that I try to run from java is supposed to create PDFs with charts and does so when executed from R itself. However, when I run the code above, nothing happens. Do you have any clues on what might be the problem?
have you tried the R_source() method?
ReplyDeleteit was newly updated in google code.
For example:
caller.R_source("/path/to/myfile.r")
hi.. i'm running a script and i'm also using R_source() method. how can i retrieve data from the script. I want to recover a variable, or some other object
DeleteHi, I tried to use Rcall to run a regression, but it returns this error:
ReplyDelete[Fatal Error] Routput4578110286621389122:1:1: Premature end of file.
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException: Premature end of file.
Anyone know anything?
Post your code
ReplyDeleteHey, we use RCaller a lot and it is totally amazing. So thanks a lot for this.
ReplyDeleteIn one of our "big" calculations, we make multiple calls to R. And by multiple, I mean about 6000. Is there some need to close these calls everytime?
Our code is currently failing because of some exception :"Too many files open. Java Exception :24". This might not even be a problem with R but we just wanted to be sure. Thanks a lot for RCaller again though.
many Thanks.
ReplyDeleteI am not sure for what the reason you have "too many open file" errors when running RCaller but in many situtations it is about the operating system level. Maybe you can search the ways for re-configurate your operating system. Or send me the source code that throws the exception.
I was getting that same "Premature end of file error" on windows XP using Eclipse when I tried to run the included Example[1,6] files. I fixed the error by opening an instance of R and running the following:
ReplyDelete> install.packages("Runiversal")
...With this package installed, I am now able to run the included Example[1,6] files.
Hi ,
ReplyDeleteI am using Windows 7 , Eclipse IDE , program is running but i am getting an empty output . Here is the code:
import java.io.File;
import javax.swing.ImageIcon;
import rcaller.RCaller;
public class Test1 {
public static void main(String[] args) {
Test1 a=new Test1();
}
/*
* Test for simple plots
*/
// @SuppressWarnings("deprecation")
public Test1() {
try {
RCaller caller = new RCaller();
caller.R_source("C:/Program Files/R/R-2.13.2/bin/x64/Rscript");
caller.cleanRCode();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
caller.addDoubleArray("x", numbers);
File file = caller.startPlot();
caller.addRCode("plot.ts(x)");
caller.endPlot();
//caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
System.out.println("success");
} catch (Exception e) {
System.out.println(e.toString());
}
}
}
Thank you! You have just discovered a bug!
ReplyDeleteI have fixed it and pushed it with new enhancements as RCaller-2.0.7.jar.
You can change your source as below:
public class Example7 {
public static void main(String[] args) {
new Example7();
}
public Example7() {
try {
RCaller caller = new RCaller();
/*
* This must be caller.setRScriptExecutable() instead.
*/
caller.setRscriptExecutable("C:/Program Files/R/R-2.13.2/bin/x64/Rscript");
/*
* After version 2.0.6
* Graphics themes change the theme for plots.
* Themes:
* BlackTheme
* SkyTheme
* DefaultTheme
*/
caller.setGraphicsTheme(new BlackTheme());
/*
* After version 2.0.6
* We build the code using RCode class. Older
* methods are deprecated and will no longer will be
* available in RCaller 3.0
*/
RCode code = new RCode();
code.clear();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
code.addDoubleArray("x", numbers);
File file = code.startPlot();
code.addRCode("plot.ts(x)");
code.endPlot();
/*
* After version 2.0.6
* We set the code.
*/
caller.setRCode(code);
caller.runOnly();
ImageIcon ii = code.getPlot(file);
code.showPlot(file);
System.out.println("success");
} catch (Exception e) {
System.out.println(e.toString());
}
}
}
This code includes some new functionality of RCaller, which is candidate for version 3.
Best.
Awesome, helped solve all of the deprecation warning... worked like a charm for a number of examples facing the same issue...
Deletehas anyone tried using RCaller to import a chart in a jsp page?
ReplyDeleteEverything works fantastically by the way, the too many open files was an issue with another part of the java code :)
Hi,
ReplyDeleteI am writing a sample code to normalise a microarray data using the loess normalisation code. Please find the code along side:
import java.io.File;
import java.lang.*;
import java.util.*;
import java.awt.image.DataBuffer;
import rcaller.RCaller;
public class Example1 {
public static void main(String[] args) {
try{
String data;
data="C:/Project_WRAIR/US09493743_251527910706_1_1 T10-105_5day_24hrs";
RCaller caller = new RCaller();
caller.cleanRCode();
caller.setRscriptExecutable("C:/Program Files/R/R-2.13.1/bin/x64/Rscript");
caller.addRCode("library(limma)");
caller.addRCode("RG <- read.maimages(tar, columns = list(G = 'gMedianSignal', Gb = 'gBGMedianSignal', R = 'rMedianSignal',Rb = 'rBGMedianSignal'), annotation = c('Row', 'Col','FeatureNum', 'ControlType','ProbeName'))");
caller.addRCode("RG <- backgroundCorrect(RG, method='minimum', offset=1)");
caller.addRCode("MA <- normalizeWithinArrays(RG, method='loess')");
caller.addRCode("A<-MA$A");
caller.runAndReturnResult("A");
double[] A=caller.getParser().getAsDoubleArray("A");
System.out.println("A");
for (int i=0;i<A.length;i++ ){
System.out.println(i+" = "+A[i]);
}
System.out.println("Bye.");
}
catch (Exception e){
System.out.println(e.toString());
}
}
}
I am getting the following error message:
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException; systemId: file:/C:/Users/SURO_B~1/AppData/Local/Temp/Routput6849780398619001039; lineNumber: 1; columnNumber: 1; Premature end of file.
Can you suggest what is going wrong? I can send you the input file if you want
2 typo errors:
ReplyDelete1.data="C:/Project_WRAIR/US09493743_251527910706_1_1 T10-105_5day_24hrs.txt" instead of
data="C:/Project_WRAIR/US09493743_251527910706_1_1 T10-105_5day_24hrs"
2.RG <- read.maimages(data, columns = list(G = 'gMedianSignal', Gb = 'gBGMedianSignal', R = 'rMedianSignal',Rb = 'rBGMedianSignal'), annotation = c('Row', 'Col','FeatureNum', 'ControlType','ProbeName')) instaed of
RG <- read.maimages(tar, columns = list(G = 'gMedianSignal', Gb = 'gBGMedianSignal', R = 'rMedianSignal',Rb = 'rBGMedianSignal'), annotation = c('Row', 'Col','FeatureNum', 'ControlType','ProbeName'))
for "too many files" error:
ReplyDeletedid you try to delete the files in temp directory with prefix "rcaller*" and "Routput*" periodically? Because RCaller saves the generated R code and XML output in temp directory with those patterns.
Is there a way to do that automatically using the caller object? Once you are done with the R code and you got your data, you don't really need these files anymore.
DeleteThat would be really useful.
We will implement some code to remove temporary codes. Indeed, there are some improvements on version 2.2.
DeleteHave a look at the page http://stdioe.blogspot.com.tr/2014/04/rcaller-220-is-released.html
RCallerPhp is now ready for testing and trying.
ReplyDeleteSee the blog entry at http://stdioe.blogspot.com/2011/10/rcallerphp-is-ready-for-testing.html
Hi,
ReplyDeleteI have problems when porting my java code from local Linux machine to Windows 2008 server. On Linux RCaller works like a charm, but on Windows Rscript.exe isn't called at all. I suspect it is a problem either:
- path notation of Rscript.exe (\\ or /)
- 8.3 length path names
- location of tmp directory
Are any known issues on Windows, regarding this area? I am using RCaller 2.0.7.
Regards,
Sašo Celarc
Hi,
ReplyDeleteI am using win7 64bit and Rcaller-2.0.7.jar.
I installed Runiversal package in R, and was able to run your sample code (e.g. test3()).
However, I had problem with my own code: possibly due to the library I am using. The library is e1071.
Here is the error message:
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException: Premature end of file.
Here is the code: If I use caller.runOnly(), it terminated successfully.
public class Rsvm {
public static void test(){
//============================
Random gen = new Random();
double x[] = new double [10];
double y[] = new double [10];
for(int i = 0 ; i < x.length ; i++){
if ( i % 2 == 0 ){
x[i] = gen.nextDouble();
y[i] = 1;
}
else{
x[i] = gen.nextDouble()*(-1.0);
y[i] = -1;
}
}
//=====================
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:\\Program Files\\R\\R-2.13.2\\bin\\Rscript.exe");
caller.addRCode("library(e1071)");
//=====================
caller.addDoubleArray("data", x);
caller.addDoubleArray("label", y);
caller.addRCode("model1 <- svm(data1,classes1,type=\'C\',kernel=\'linear\')");
caller.addRCode("z1=predict(model1,1.34)");
caller.addRCode("z=as.numeric(z1)");
caller.runAndReturnResult("z");
}
public static void main(String [] args){
test();
}
}
hello there,
ReplyDeleteWe're trying example7 in eclipse running on windows 7.
But when the plot is generated it appears empty, and it shows no exceptions or errors.
When we looked up the generated plot it has a size of 0 kb.
Code:
import java.io.File;
import javax.swing.ImageIcon;
import graphics.BlackTheme;
import rcaller.RCaller;
import rcaller.RCode;
public class Plotmachine {
public static void main(String[] args) {
new Plotmachine();
}
public Plotmachine() {
try {
RCaller caller = new RCaller();
/*
* This must be caller.setRScriptExecutable() instead.
*/
caller.setRscriptExecutable("D:/Programma's/R/R-2.12.2/bin/R.exe");
/*
* After version 2.0.6
* Graphics themes change the theme for plots.
* Themes:
* BlackTheme
* SkyTheme
* DefaultTheme
*/
caller.setGraphicsTheme(new BlackTheme());
/*
* After version 2.0.6
* We build the code using RCode class. Older
* methods are deprecated and will no longer will be
* available in RCaller 3.0
*/
RCode code = new RCode();
code.clear();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
code.addDoubleArray("x", numbers);
File file = code.startPlot();
code.addRCode("plot.ts(x)");
code.endPlot();
/*
* After version 2.0.6
* We set the code.
*/
caller.setRCode(code);
caller.runOnly();
ImageIcon ii = code.getPlot(file);
System.out.println(ii.toString());
code.showPlot(file);
System.out.println("success");
} catch (Exception e) {
System.out.println(e.toString());
}
}
}
Have you your Runiversal package installed in R after the R 2.14 update?
ReplyDeleteI just installed it but it still doesn't work.
ReplyDeleteNL:
ja mejong, maar t werkt nog steeds niet :/
by the version 2.2, you no longer need to install Runiversal package. have a look at the page http://stdioe.blogspot.com.tr/2014/04/rcaller-220-is-released.html
DeleteThe path you have written "D:/Programma's/R/R-2.12.2/bin/R.exe" should be "D:\\Programma's\\R/R-2.12.2\\bin\\R.exe" because you are in a Windows System. Secondly, the Rscript executable file is "Rscript.exe" in windows so the final path should be D:\\Programma's\\R\\R-2.12.2\\bin\\Rscript.exe.
ReplyDeleteI switch between my school computer (which runs Linux) and my home computer every day and I always make this mistake :( Thanks a lot for noticing! It runs perfectly now.
ReplyDeleteThis is a hint for all those struggling with "premature end of file" errors.
ReplyDeleteThe best way to debug those is to find the rcaller input file (here [winxp] it is in the locale settings, user name, local application data, temp folder) and try to run this rcaller123456... file via CLI call of Rscript.exe
Hi, me again. After solving my “premature end of file” issue, I’m faced with an other problem. Reason is probably me first calling rCaller.runOnly(), then calling rCaller.runAndReturnResult() without proper handling of the rCaller inbetween. Here is my code, any help is appreciated!
ReplyDelete// Setup
RCaller rCaller = new RCaller();
rCaller.setRscriptExecutable("C:\\Dokumente und Einstellungen\\mzuba\\Lokale Einstellungen\\Temp\\Programme\\R-2.13.0\\bin\\Rscript.exe");
rCaller.cleanRCode();
// Read and prepare data
rCaller.R_source("Z:\\\\Novell\\\\eclipse-workspace\\\\ltcmicroR\\\\ShareMergedForCalculations.R");
rCaller.runOnly();
[…]
rCaller.cleanRCode();
rCaller.addRCode("mylogit <- glm(w2health.cond.${cond.getShareName()} ~ ${coefList}, ${subsetCondition}, family=binomial(\"logit\"))");
rCaller.runAndReturnResult("mylogit");
// JRE crashes here and says “Can not run C:\Dokumente und Einstellungen\mzuba\Lokale Einstellungen\Temp\Programme\R-2.13.0\bin\Rscript.exe. Reason: java.lang.IllegalThreadStateException”
[…]
}
Hello,
ReplyDeleteI'm having trouble changing the ouput location for the graph I'm trying to make.
I'm still using example code and tried moving the file with plain java with this:
File source = ....
File destination = ...
if (!destination.exists())
{
source.renameTo(destination);
}
As well as using code.addRcode("setwd(\" \")").
This is the code I'm using.
import java.io.File;
import javax.swing.ImageIcon;
import graphics.*;
import rcaller.RCaller;
import rcaller.RCode;
public {
try {
RCaller caller = new RCaller();
/*
* This must be caller.setRScriptExecutable() instead.
*/
caller.setRscriptExecutable("/usr/bin/Rscript");
/*
* After version 2.0.6
* Graphics themes change the theme for plots.
* Themes:
* BlackTheme
* SkyTheme
* DefaultTheme
*/
caller.setGraphicsTheme(new DefaultTheme());
/*
* After version 2.0.6
* We build the code using RCode class. Older
* methods are deprecated and will no longer will be
* available in RCaller 3.0
*/
RCode code = new RCode();
code.clear();
code.addRCode("setwd(\"\")");
double[] numbers = new double[]{1, 4, 3, 5, 6, 9, 19};
code.addDoubleArray("y", numbers);
File file = code.startPlot();
code.addRCode("barplot(sub = \"this chart shows the amount of enzymes per corresonding EC class\",main = \"Amount of enzymes per class\" ,ylab = \"Amount in file\", xlab= \"EC Classes\",y,beside= true, col = rainbow(7), legend.text=c(\"EC class 1\",\"EC class 2\",\"EC class 3\",\"EC class 4\",\"EC class 5\",\"EC class 6\",\"EC class 7\"), args.legend =list(x= \"topleft\"))");
code.endPlot();
/*
* After version 2.0.6
* We set the code.
*/
caller.setRCode(code);
caller.runOnly();
ImageIcon ii = code.getPlot(file);
System.out.println(ii.toString());
code.showPlot(file);
System.out.println("success");
} catch (Exception e) {
System.out.println(e.toString());
}
}
}
PS:
Is there any java doc available I only found the lose files not the whole package.
I do linear regression
ReplyDeleteCall:
lm(formula = y ~ x + I(x^2))
Residuals:
1 2 3 4 5
0.2571 -0.6286 0.3429 0.1714 -0.1429
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 2.483e-15 1.202e+00 0.000 1.000
x 6.714e-01 9.163e-01 0.733 0.540
I(x^2) 7.143e-02 1.498e-01 0.477 0.681
Residual standard error: 0.5606 on 2 degrees of freedom
Multiple R-squared: 0.9509, Adjusted R-squared: 0.9018
F-statistic: 19.36 on 2 and 2 DF, p-value: 0.04911
When i run it through RCaller i can't get back the last p-value (near F-statistic).
Available results from lm() object:
[call, terms, residuals, coefficients, aliased, sigma, df, r_squared, adj_r_squared, fstatistic, cov_unscaled]
It seems that this one is missing? or i can obtain it somewhat differently?
since lm object does not include the p-value, you have to use the anova() function. Simply
Deleteols <- lm(formula = y ~ x + I(x^2))
a <- anova(ols)
pval <- a["Pr(>F)"]
gets the p-value in variable pval. Then, you can combine the results in a single variable:
ols["pvalue"] <- pval
After all, your list of available objects becomes:
[call, terms, residuals, coefficients, aliased, sigma, df, r_squared, adj_r_squared, fstatistic, cov_unscaled, pvalue]
Hope this solves.
This comment has been removed by the author.
ReplyDeletehow can I handle an error? I mean i run regression for example but for some reason (wrong initial values or whatever) is not working.. how can i handle that?
ReplyDeletecaller.addRCode("exponentialmodel = nls(y ~ exp(x * a))");
caller.runAndReturnResult("exponentialmodel");
Could you provide an example on how to handle such case? We may need to use an R command? that will return back an empty/null object in either case so we can check it??
thx
you can try the r side exception & conditional handling
ReplyDeleteas described at http://stat.ethz.ch/R-manual/R-patched/library/base/html/conditions.html
I called any methods and they had errors :
ReplyDeleteThe method startPlot() from the type RCaller is deprecated
Help me !
Yes it is deprecated after version 2.0 but the library still supports plotting functionality. See the examples in source code repository.
Deletei wan to plot a graph, there are no error but the frame is not plotting with the graph, just a empty frame. My entire code is
ReplyDeletecaller.addRCode("library(kernlab)");
caller.addRCode("n<-50");
caller.addRCode("f1blueclass <- rnorm(n, mean = 1, sd = 0.4)");
caller.addRCode("f1redclass <- rnorm(n, mean = 1.5, sd = 0.4)");
caller.addRCode("f2blueclass <- rnorm(n, mean = 1, sd = 0.4)");
caller.addRCode("f2redclass <- rnorm(n, mean = 1.5, sd = 0.4)");
caller.addRCode(" x <- rbind(blueclass, redclass");
caller.addRCode("y <- matrix(c(rep(1,50),rep(-1,50)))");
File file = caller.startPlot();
caller.addRCode("plot(x,col=ifelse(y>0,\"blue\",\"red\")) ");
caller.addRCode("legend(\"topleft\",c('Blue Class','Red Class'),col=c(\"blue\",\"red\"),pch=1,text.col=c(\"blue\",\"red\"))");
caller.endPlot();
caller.runOnly();
caller.showPlot(file);
Hello,
ReplyDeleteI got the same error like others (with SAX).
That's my code:
int d[] = new int[] {1,0,1,0,1,1,0,1,0,1,0,0,1,0,1,1,0,0,0,1,1,1,0,0,1,0,0,0,1,1,1,1,0,0,1,1};
RCaller rc = new RCaller();
rc.setRscriptExecutable("/usr/bin/Rscript");
rc.cleanRCode();
rc.addRCode("x <- as.dist(matrix(c("+implode(d,",")+"), ncol=6, byrow=TRUE))");
rc.addRCode("hc <- hclust(x, 'complete')");
rc.addRCode("cutted <- cutree(hc, h=0)");
rc.addRCode("my.all<-list(clustered=cutted)");
rc.runAndReturnResult("my.all");
System.out.println(rc.getParser().getNames());
and error:
cat(makexml(obj=my.all, name="my.all"), file="/var/folders/1l/sby4rxgs6nn_j1hvjv7mktm00000gn/T/Routput4116407920481509008")
Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException: Premature end of file.
Could you tell me what is wrong?
Thanks in advance!
I found the error!
ReplyDeleteFor others:
You can check errors if you open folder /var/folders/1l/sby4rxgs6nn_j1hvjv7mktm00000gn/T/ , then open file called rcaller{Here some numbers}, copy code and paste it in R. Tadam you got error ;)
For me it was that makexml not exist or sth similar and that I did not have Runiversal package so...
Bottom line if you are using Mac you should manually install the Runiversal package...
At the end I have one question: How can i pass 2d array from Java to R? Right now I'm transforming 2d to vector, passing it to R and then creating matrix using information about nrows...
Hi, thanks for Rcaller, saved me a lot of time trying to get JRI/rJava to work properly!
ReplyDeleteHowever, it might be helpful to others if you made a note in the main post about getting round the 'deprecated' methods so eg:
RCaller caller = new RCaller();
caller.setRscriptExecutable("/usr/bin/Rscript");
double[] x = new double[]{1,2,3,4,5,6,7,8,9,10};
double[] y = new double[]{2,4,6,8,10,12,14,16,18,30};
caller.addDoubleArray("x", x);
caller.addDoubleArray("y", y);
caller.addRCode("ols<-lm(y~x)");
should become:
RCaller caller = new RCaller();
caller.setRscriptExecutable("/usr/bin/Rscript");
RCode code = new RCode();
//double[]s as before...
code.addDoubleArray("y", y); //etc...
code.addRCode("ols<-lm(y~x)");
caller.setRCode(code);
Thanks again!
Hi,
DeleteWe are planning to publish more documentation with RCaller in its next release.
Thank you for your concern.
Hi,
ReplyDeleteSeems I'm running into a familiar problem:
Exception in thread "main" rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException: Premature end of file.
Additionally, I get a bunch of other messages:
[Fatal Error] Routput3105202150200792412:1:1: Premature end of file.
packageExist<-require(Runiversal)
if(!packageExist){
install.packages("Runiversal", repos=" http://cran.r-project.org")
}
scores<-c(15.49, ... , 14.97);
classes<-c(1, ... , 1);
library(ROCR)
pred <- prediction(scores,classes)
auc <- as.numeric(performance(pred,'auc')@y.values)
cat(makexml(obj=auc, name="auc"), file="C:/DOCUME~1/Ali/LOCALS~1/Temp/Routput3105202150200792412")
This is the relevant code:
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:\\Program Files\\R\\R-2.15.0\\bin\\Rscript.exe");
RCode code = new RCode();
code.clear();
code.addDoubleArray("scores", r_scores);
code.addIntArray("classes", r_classes);
code.addRCode("library(ROCR)"); //add rocr library
code.addRCode("pred <- prediction(scores,classes)");
code.addRCode("auc <- as.numeric(performance(pred,'auc')@y.values)");
caller.setRCode(code);
caller.runAndReturnResult("auc");
//double auc = caller.getParser().getAsDoubleArray("auc")[0];
System.out.println(caller.getParser().getNames());
//System.out.println("ROC AUC: " + auc);
I want to run this code for a large number of datasets (say 1000) and this works fine for 3, but crashes on the 4th. The file that the error points to is blank. I updated to R 2.15.0 and have installed Runiversal so not quite sure where to go from here - any ideas would be appreciated. Thanks in advance!
Ok, it seems my problem is not with Rcaller, but the external library I'm using, will post again once I've got it sorted out...
DeleteHello, I'm using your solution with apache tomcat 6.0 and it runs all right, but generates a. PNG TEMP void in tomcat and a file with the script. If I run this script directly on the graph it generates R (. PNG) normally. What can be?
ReplyDeletei tried to run the code in the link but it returns an empty console without any graph. Even the .PNG file is empty. I am using windows.
ReplyDeleteThis is an awesome project well done! Is there a way to run complete R scripts in one go? i.e. rcaller.setRCode("/path/to/file.r");
ReplyDeleteThis project is amazing! But I do have one problem when I try to deal with large data set. For example, if I want to use linear regression and have a large matrix (50000 * 10) as one of the inputs, I am doomed. The Rcaller will try to generate the matrix in script and send it to R (I knew this from the script generated in C:\Users\XXX\AppData\Local\Temp), which takes a long time... while in R it is much much faster.
ReplyDeleteThe code looks like this:
code.addDoubleMatrix("data.train", featureTrain);
code.addDoubleMatrix("data.test", featureTest);
code.addDoubleArray("upTrend", upTrend);
code.addRCode("allTrainData = cbind(upTrend, data.train)");
code.R_source("Here is my source file");
code.addRCode("res = function(inputs)");
caller.setRCode(code);
caller.runAndReturnResult("res$pred");
pred = caller.getParser().getAsDoubleArray("res$pred");
The same code runs very fast in R. Thanks a lot!
Hello,
DeleteI try to use an .Rdata file but it doesn't work. Actually I don't know if the issue is this?
I've tried several solutions like declaring and loading this file:
String file = "/Users/bendarraz/Documents/ObjTable.RData";
code.addRCode("load(file)");
but I've this warning:"The value of the local variable file is not used".
Other solution: caller.R_source("/Users/bendarraz/Documents/ObjTable.RData");
But still wrong.
Here is my R code
> file.choose()
[1] "/Users/bendarraz/Documents/ObjTable.RData"
> file<-file.choose()
> load(file)
> seed.landslide <- sample(ObjTable$Mean.OID[(which(ObjTable$class_name == 'landslide'))], 1)
> seed.non.landslide <- sample(ObjTable$Mean.OID[(which(ObjTable$class_name == 'non landslide'))], 1)
> selection <- c(seed.landslide, seed.non.landslide)
> selection
[1] 56736 105804
> new.OID <- query(ObjTable, selection, n.trees=50)
> new.OID
[1] 301848
and my Java code:
caller.setRscriptExecutable("/usr/bin/Rscript");
String file = "/Users/bendarraz/Documents/ObjTable.RData";
/*
* Adding R Code
*/
code.addRCode("load(file)");
code.addRCode("seed.landslide <- sample(ObjTable$Mean.OID[(which(ObjTable$class_name == 'landslide'))], 1)");
code.addRCode("seed.non.landslide <- sample(ObjTable$Mean.OID[(which(ObjTable$class_name == 'non landslide'))], 1)");
code.addRCode("selection <- c(seed.landslide, seed.non.landslide)");
code.addRCode("toto <- list(selection)");
caller.setRCode(code);
caller.runAndReturnResult("toto");
double[] results;
results = caller.getParser().getAsDoubleArray("selection");
System.out.println("results " + results[0]);
code.addRCode ("source(/Users/bendarraz/Documents/query.r)");
code.addRCode("nouveau.OID <- query(ObjTable, selection, n.trees=50)");
double[] results;
results = caller.getParser().getAsDoubleArray("nouveau.OID");
System.out.println("nouveau.OID" + results[0]);
thank you in advance.
This is a great find, thank you!!
ReplyDeleteI'm using RCaller to generate PNGs from Hadoop.
Alan
nice!
DeleteHello,
ReplyDeleteI am launching rcaller within a service and when there is an error in the script I am using, instead of throwing back the error, it just stop answering and the whole service is blocked. Is there any function that would remove this?
thanks
A guy uploaded a video on RCaller in Youtube. Thanks for the uploader.
ReplyDeletehttp://stdioe.blogspot.com/2012/08/a-nice-video-on-rcaller-20.html
I want to plot a graph, using qcc, there are no error but the frame is not plotting with the graph, just a empty frame.
DeleteI am runnning the following code in Netbeans, using Windows 7 (64 bits), RCaller version in 2.0.7 and R version in 2.15.1
My entire code is:
import rcaller.RCaller;
import java.io.File;
public class TestRCaller1{
public void test1(){
try{
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:/Program Files/R/R-2.15.1/bin/x64/Rscript");
caller.cleanRCode();
caller.addRCode("library(MASS)");
caller.addRCode("library(qcc)");
caller.addRCode("require(qcc)");
caller.addRCode("Teso3<-read.csv('C:/Users/dell/Documents/Disconformteso3trim.csv',header=T)");
caller.addRCode("attach(Teso3)");
caller.addRCode("Disconform9<-qcc.groups(Disconform9,Sample3)");
caller.addRCode("tmuestra<-scan(C:/Users/dell/Documents/tmuestra.csv,sep=';')");
// //caller.addRCode("pdf('grafico.pdf')");
File file = caller.startPlot();
caller.addRCode("grafico<-qcc(Disconform9[1:65],type='p',sizes=tmuestra)");
caller.addRCode("plot.ts('grafico')");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
}catch(Exception e){
System.out.println(e.toString());}
}
Can you help me to see where the problem is?
Thanks.
Jess
Hi,
ReplyDeleteI am calling a function in "outliers" packages. I installed this package in my r interpreter. but I got error when I test it in java. Here is my code and out put.
Code:
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:\\Program Files\\R\\R-2.15.1\\bin\\Rscript.exe");
RCode code= new RCode();
code.clear();
code.addRCode("x <-0.9");
code.addRCode("n <- 3");
code.addRCode("g <- qgrubbs(x,n)");
caller.setRCode(code);
caller.runAndReturnResult("g");
I got this error:
rcaller.exception.ParseException: Can not handle R results due to : rcaller.exception.ParseException: Can not parse output: The generated file C:\Users\I833891\AppData\Local\Temp\2\Routput4760557801250804811 is empty
I don't see why it returns an empty file. Do you support the functions in outliers package?
Thanks
I want to plot a graph, using qcc, there are no error but the frame is not plotting with the graph, just a empty frame.
DeleteI am runnning the following code in Netbeans, using Windows 7 (64 bits), RCaller version in 2.0.7 and R version in 2.15.1
My entire code is:
import rcaller.RCaller;
import java.io.File;
public class TestRCaller1{
public void test1(){
try{
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:/Program Files/R/R-2.15.1/bin/x64/Rscript");
caller.cleanRCode();
caller.addRCode("library(MASS)");
caller.addRCode("library(qcc)");
caller.addRCode("require(qcc)");
caller.addRCode("Teso3<-read.csv('C:/Users/dell/Documents/Disconformteso3trim.csv',header=T)");
caller.addRCode("attach(Teso3)");
caller.addRCode("Disconform9<-qcc.groups(Disconform9,Sample3)");
caller.addRCode("tmuestra<-scan(C:/Users/dell/Documents/tmuestra.csv,sep=';')");
// //caller.addRCode("pdf('grafico.pdf')");
File file = caller.startPlot();
caller.addRCode("grafico<-qcc(Disconform9[1:65],type='p',sizes=tmuestra)");
caller.addRCode("plot.ts('grafico')");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
}catch(Exception e){
System.out.println(e.toString());}
}
Can you help me to see where the problem is?
Thanks.
Jess
I want to plot a graph, using qcc, there are no error but the frame is not plotting with the graph, just a empty frame.
ReplyDeleteI am runnning the following code in Netbeans, using Windows 7 (64 bits), RCaller version in 2.0.7 and R version in 2.15.1
My entire code is:
import rcaller.RCaller;
import java.io.File;
public class TestRCaller1{
public void test1(){
try{
RCaller caller = new RCaller();
caller.setRscriptExecutable("C:/Program Files/R/R-2.15.1/bin/x64/Rscript");
caller.cleanRCode();
caller.addRCode("library(MASS)");
caller.addRCode("library(qcc)");
caller.addRCode("require(qcc)");
caller.addRCode("Teso3<-read.csv('C:/Users/dell/Documents/Disconformteso3trim.csv',header=T)");
caller.addRCode("attach(Teso3)");
caller.addRCode("Disconform9<-qcc.groups(Disconform9,Sample3)");
caller.addRCode("tmuestra<-scan(C:/Users/dell/Documents/tmuestra.csv,sep=';')");
// //caller.addRCode("pdf('grafico.pdf')");
File file = caller.startPlot();
caller.addRCode("grafico<-qcc(Disconform9[1:65],type='p',sizes=tmuestra)");
caller.addRCode("plot.ts('grafico')");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
}catch(Exception e){
System.out.println(e.toString());}
}
I am trying to call an R script from my java code. Here is the R script:
ReplyDeletetest <- list()
test[[1]] <- c(1:10)
test[[2]] <- c("hallo", "welt", "ich", "bin", "da")
test[[3]] <- matrix("NA",ncol=2,nrow=15)
names(test) <- c("number", "chars", "matrix")
bla <- "blubl bulb bulbu"
return(list(bla, test))
Now here comes my Java code:
public static void main(String[] args) {
// TODO Auto-generated method stub
String x = null;
RCaller caller = new RCaller();
caller.setRscriptExecutable("/home/bio/groupshare/software/R-2.15.1/bin/Rscript");
caller.cleanRCode();
caller.R_source("/RBIqualityAssessmentRB/trunk/resources/rScripts/test.R");
caller.runAndReturnResult(x);
System.out.println(x);
}
There are errors everytime i change something. Could you please tell me where is the problem.
hello
ReplyDeletei have two problems the first onr is not very important i think:
rcaller.addDoubleArray("x", numbers);
File file = rcaller.startPlot();
rcaller.addRCode("plot(x)");
rcaller.endPlot();
rcaller.runOnly();
ImageIcon ii = rcaller.getPlot(file);
rcaller.showPlot(file);
all this methods are deprecated ????
and for the method:
RCaller rcaller = new RCaller();
rcaller.setRExecutable("C:/Documents and Settings/My Documents/R/R-2.15.1/bin/Rscript");
i tried all the possiblities for the Rscript.exe but i have always the error:
Exception in thread "main" rcaller.exception.RCallerExecutionException: RscriptExecutable is not defined. Please set this variable to full path of Rscript executable binary file.
at rcaller.RCaller.runOnly(RCaller.java:282)
at Rcaller.main(Rcaller.java:27)
and i dont know way, if someone has an idea??
thank's
try rcaller.setRScriptExecutable().
DeleteYes they are deprecated. Use RCode to generate code and then attach it to Rcaller using rcaller.setRCode(code).
RCaller caller = new RCaller();
ReplyDeleteRCode rCode = new RCode();
caller.setRscriptExecutable("/usr/local/lib64/R/bin/Rscript");
caller.cleanRCode();
rCode.addRCode("library(Runiversal)");
rCode.addRCode("library(sentiment)");
rCode.addRCode("library(DBI)");
rCode.addRCode("library(RMySQL)");
rCode.addRCode("library(rJava)");
rCode.addRCode("library(rhdfs)");
rCode.addRCode("hdfs.init()");
rCode.addRCode("totData<-readLines(\"/home/madamanchik/Desktop/write.txt\")");
rCode.addRCode("tweetData=\"\"");
caller.setRCode(rCode);
caller.runAndReturnResult("totData");
I am getting the below error
ReplyDelete[Fatal Error] Routput4458404778467616229:1:1: Premature end of file.
cat(makexml(obj=totData, name="totData"), file="/tmp/Routput4458404778467616229")
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException; systemId: file:/tmp/Routput4458404778467616229; lineNumber: 1; columnNumber: 1; Premature end of file.
Please suggest but i have already installed Runiversal package in R
This issue is resolved..... :)
ReplyDeleteHow did U resolved it?
DeleteHello
ReplyDeleteI'm trying RCaller in order to execute BG-NBD analysis
I downloaded this jar "RCaller-2.1.1-SNAPSHOT.jar" and I tried to execute the R code listed on this url: http://code.google.com/p/clv-master-thesis/. I need to use this R code: http://code.google.com/p/clv-master-thesis/source/browse/trunk/model-bg-nbd.R
Now I installed the Runiversal package and I tried this sample:
RCaller caller = new RCaller();
caller.setRscriptExecutable("/usr/bin/Rscript");
RCode code = new RCode();
code.clear();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
code.addDoubleArray("x", numbers);
File file = code.startPlot();
System.out.println("Plot will be saved to : " + file);
code.addRCode("plot.ts(x)");
code.endPlot();
caller.setRCode(code);
caller.runOnly();
code.showPlot(file);
All works pretty good
I wanted to adapt it to my scenario and I wrote this unit test:
InputStreamReader reader = new InputStreamReader(Thread.currentThread().getContextClassLoader().getResourceAsStream("model-bg-nbd.R"));
BufferedReader br = new BufferedReader(reader);
String line = null;
RCaller caller = new RCaller();
caller.setRscriptExecutable("/usr/bin/Rscript");
caller.cleanRCode();
RCode code = new RCode();
code.clear();
int dimensione = 5;
List datas = new ArrayList(dimensione);
String dataFrameElement = null;
for(int i = 0; i < dimensione; i++){
if( i == 0 ){
dataFrameElement = "ID Tempo Utente Test";
}else{
dataFrameElement = Math.random()*100+" "+Math.random()*100+" "+Math.random()*100+" "+Math.random()*100;
}
datas.add(dataFrameElement);
}
code.addStringArray("data", datas.toArray(new String[datas.size()]));
while( (line = br.readLine()) != null ){
if(!line.startsWith("#")){
code.addRCode(line);
}
}
String doMle = "mle<-bgEstimateParameters(data, list(r=1, alpha=2, a=1, b=2));";
code.addRCode(doMle);
caller.setRCode(code);
caller.runAndReturnResult("mle");
String[] results = caller.getParser().getAsStringArray("mle");
logger.info(results[0]);
} catch (Exception e) {
logger.error(e.getMessage(), e);
}
When I execute it I get this error:
.
.
.
.
mle<-bgEstimateParameters(data, list(r=1, alpha=2, a=1, b=2));
cat(makexml(obj=mle, name="mle"), file="/tmp/Routput359342344387748448")
13:30:23,228 ERROR [TestScriptingLanguagesCalling] Can not handle R results due to : rcaller.exception.ParseException: Can not parse output: The generated file /tmp/Routput359342344387748448 is empty
rcaller.exception.ParseException: Can not handle R results due to : rcaller.exception.ParseException: Can not parse output: The generated file /tmp/Routput359342344387748448 is empty
at rcaller.RCaller.runAndReturnResult(RCaller.java:412)
I can't figure what I'm wrong.
Can you give me any tips?
Cheers,
Angelo
Hai Everyone,
ReplyDeleteI am just tring to read protein sequences from online DB. It works well in R but when i try to execute via Java using RCaller i get the following error:
[Fatal Error] Routput34681:1:1: Premature end of file.
packageExist<-require(Runiversal)
if(!packageExist){
install.packages("Runiversal", repos=" http://cran.r-project.org")
}
library("seqinr")
retrieveseqs <- function(seqnames,acnucdb)
{
myseqs <- list()
require("seqinr")
choosebank(acnucdb)
for (i in 1:length(seqnames))
{
seqname <- seqnames[i]
print(paste("Retrieving sequence",seqname,"..."))
queryname <- "query2"
query <- paste("AC=",seqname,sep="")
query(`queryname`,`query`)
seq <- getSequence(query2$req[[1]])
myseqs[[i]] <- seq
}
closebank()
return(myseqs)
}
seqnames <- c("ACY70669","ACQ44515","Q6YMS4","Q58HT7")
seqs <- retrieveseqs(seqnames,"swissprot")
cat(makexml(obj=seqs, name="seqs"), file="C:/Users/PRINCI~1/AppData/Local/Temp/Routput34681") // this file is Empty
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException: Premature end of file.
Please Help.....
Hai Everyone,
ReplyDeleteI already posted about my error:[Fatal Error] Routput34681:1:1: Premature end of file.
Here is my code:
import rcaller.RCaller;
import rcaller.RCode;
public class ReadSeq
{
public ReadSeq()
{
try
{
RCaller caller = new RCaller();
RCode code=new RCode();
code.clear();
caller.setRscriptExecutable("C:\\Program Files\\Option\\R-2.15.1\\bin\\RScript.exe");
code.addRCode("library(\"seqinr\")");
code.addRCode("retrieveseqs <- function(seqnames,acnucdb)");
code.addRCode("{");
code.addRCode("myseqs <- list()");
code.addRCode("require(\"seqinr\")");
code.addRCode("choosebank(acnucdb)");
code.addRCode("for (i in 1:length(seqnames))");
code.addRCode("{");
code.addRCode("seqname <- seqnames[i]");
code.addRCode("print(paste(\"Retrieving sequence\",seqname,\"...\"))");
code.addRCode("queryname <- \"query2\"");
code.addRCode("query <- paste(\"AC=\",seqname,sep=\"\")");
code.addRCode("query(`queryname`,`query`)");
code.addRCode("seq <- getSequence(query2$req[[1]])");
code.addRCode("myseqs[[i]] <- seq");
code.addRCode("}");
code.addRCode("closebank()");
code.addRCode("return(myseqs)");
code.addRCode(" }");
code.addRCode("seqnames <- c(\"ACY70669\",\"ACQ44515\",\"Q6YMS4\",\"Q58HT7\")");
code.addRCode("seqs <- retrieveseqs(seqnames,\"swissprot\")");
code.addRCode("write.fasta(seqs, seqnames, file=\"type.fasta\"");
caller.setRCode(code);
//caller.runOnly();
caller.runAndReturnResult("seqs");
String[] s=new String[10];
System.out.println("hai");
System.out.println(s[0]);
}
catch (Exception e){
System.out.println(e.toString());
}
}
public static void main(String args[])
{
ReadSeq R=new ReadSeq();
}
}
Seeking for a reply to solve the problem.....Thank You
package test;
ReplyDeleteimport java.util.Random;
import rcaller.RCaller;
/**
*
* @author Mehmet Hakan Satman
*/
public class Test3 {
public static void main(String[] args){
new Test3();
}
public Test3(){
try{
/*
* Creating Java's random number generator
*/
Random random = new Random();
/*
* Creating RCaller
*/
RCaller caller = new RCaller();
/*
* Full path of the Rscript. Rscript is an executable file shipped with R.
* It is something like C:\\Program File\\R\\bin.... in Windows
*/
caller.setRscriptExecutable("/usr/bin/Rscript");
/* We are creating a random data from a normal distribution
* with zero mean and unit variance with size of 100
*/
double[] data = new double[100];
for (int i=0;imean(x)");
caller.addRCode("my.var<-var(x)");
caller.addRCode("my.sd<-sd(x)");
caller.addRCode("my.min<-min(x)");
caller.addRCode("my.max<-max(x)");
caller.addRCode("my.standardized<-scale(x)");
/*
* Combining all of them in a single list() object
*/
caller.addRCode("my.all<-list(mean=my.mean, variance=my.var, sd=my.sd, min=my.min, max=my.max, std=my.standardized)");
/*
* We want to handle the list 'my.all'
*/
caller.runAndReturnResult("my.all");
double[] results;
/*
* Retrieving the 'mean' element of list 'my.all'
*/
results = caller.getParser().getAsDoubleArray("mean");
System.out.println("Mean is "+results[0]);
/*
* Retrieving the 'variance' element of list 'my.all'
*/
results = caller.getParser().getAsDoubleArray("variance");
System.out.println("Variance is "+results[0]);
/*
* Retrieving the 'sd' element of list 'my.all'
*/
results = caller.getParser().getAsDoubleArray("sd");
System.out.println("Standard deviation is "+results[0]);
/*
* Retrieving the 'min' element of list 'my.all'
*/
results = caller.getParser().getAsDoubleArray("min");
System.out.println("Minimum is "+results[0]);
/*
* Retrieving the 'max' element of list 'my.all'
*/
results = caller.getParser().getAsDoubleArray("max");
System.out.println("Maximum is "+results[0]);
/*
* Retrieving the 'std' element of list 'my.all'
*/
results = caller.getParser().getAsDoubleArray("std");
/*
* Now we are retrieving the standardized form of vector x
*/
System.out.println("Standardized x is ");
for (int i=0;i));
}
}
}
Hello everyone,
ReplyDeleteI am trying to use Rcaller to plot phylogenetic tree via java.
My code:
mport java.io.File;
import javax.swing.ImageIcon;
import graphics.*;
import rcaller.*;
public class phylo
{
public phylo()
{
try
{
RCaller caller = new RCaller();
RCode code=new RCode();
code.clear();
caller.setRscriptExecutable("C:\\Program Files\\Option\\R-2.15.1\\bin\\RScript");
code.addRCode("library(\"seqinr\")");
code.addRCode("library(\"ape\")");
code.addRCode("virusaln <- read.alignment(file = \"denguetype.phy\" , format = \"phylip\")");
code.addRCode("rootedNJtree <- function(alignment, theoutgroup, type)");
code.addRCode("{");
code.addRCode("require(\"ape\")");
code.addRCode("require(\"seqinr\")");
code.addRCode("makemytree <- function(alignmentmat, outgroup=`theoutgroup`)");
code.addRCode("{");
code.addRCode("alignment <- ape::as.alignment(alignmentmat)");
code.addRCode("if(type == \"protein\")");
code.addRCode("{");
code.addRCode("mydist <- dist.alignment(alignment)");
code.addRCode("}");
code.addRCode("else if (type == \"DNA\")");
code.addRCode("{");
code.addRCode("alignmentbin <- as.DNAbin(alignment)");
code.addRCode("mydist <- dist.dna(alignmentbin)");
code.addRCode("}");
code.addRCode("mytree <- nj(mydist)");
code.addRCode("mytree <- makeLabel(mytree, space=\"\")");
code.addRCode("myrootedtree <- root(mytree, outgroup, r=TRUE)");
code.addRCode("return(myrootedtree)");
code.addRCode("}");
code.addRCode("mymat <- as.matrix.alignment(alignment)");
code.addRCode("myrootedtree <- makemytree(mymat, outgroup=theoutgroup)");
code.addRCode("myboot <- boot.phylo(myrootedtree, mymat, makemytree)");
code.addRCode("plot.phylo(myrootedtree, type=\"p\")");
code.addRCode("nodelabels(myboot,cex=0.7)");
code.addRCode("myrootedtree$node.label <- myboot");
code.addRCode("return(myrootedtree)");
code.addRCode("}");
code.addRCode("tree1<-rootedNJtree(virusaln,\"ACY70669\",type=\"protein\")");
File file = code.startPlot();
System.out.println("Plot will be saved to : " + file);
code.addRCode("plot(tree1)");
code.endPlot();
caller.setRCode(code);
caller.runOnly();
//ImageIcon ii = code.getPlot(file);
code.showPlot(file);
}
catch (Exception e)
{
System.out.println(e.toString());
}
}
public static void main(String args[])
{
new phylo();
}
}
While i run the code in R, the tree is created automatically. But in java, i tried to plot using startplot but nothing happened and the created png file is empty. Please guide me.....Thank You
Hi Everyone
ReplyDeleteI am trying to implement clustering using R in java by employing R caller. I am trying to run sample code for clustering validation and I get that common error faced by most of the users: Premature end of file
The code is:
public test3()
{
try{
RCaller caller = new RCaller();
caller.cleanRCode();
caller.setRscriptExecutable("C:/Program Files/R/R-2.15.1/bin/x64/Rscript");
caller.addRCode("library(clvalid)");
caller.addRCode("data(mouse)");
caller.addRCode("express <- mouse [,c(M1,M2,M3,NC1,NC2,NC3)]");
caller.addRCode("rownames (express) <- mouse$ID ");
caller.addRCode("intern <- clValid(express, 2:6 , clMethods = c( hierarchical,kmeans,diana,clara,model) ,validation = internal)");
caller.addRCode("b <- summary(intern) ");
caller.runOnly();
caller.runAndReturnResult("b");
}
catch (Exception e){
e.printStackTrace();
}
}
}
The exact error is:
[Fatal Error] Routput2989518813454919276:1:1: Premature end of file.
rcaller.exception.RCallerExecutionException: Can not handle R results due to : rcaller.exception.RCallerParseException: Can not parse the R output: org.xml.sax.SAXParseException; systemId: file:/C:/Users/RAVINDER/AppData/Local/Temp/Routput2989518813454919276; lineNumber: 1; columnNumber: 1; Premature end of file.
at rcaller.RCaller.runAndReturnResult(RCaller.java:402)
at test.test3.(test3.java:29)
at test.test3.main(test3.java)
Its urgent..please respond
Thanks
Ravinder
Open the R console from command line and type the following
Deleteinstall.packages ("Runiversal")
Once this is done, try it again, it will work
hi,
ReplyDeletei am using rcaller and i have successfully executed an example pgm to output a graph. in my application i have defined a rfunction in a rscript file and the function displays a pie chart. how can i display this graph directly in java??
pls help
here's d rfunction
academicpie<-function(){
library(RPostgreSQL)
drv<-dbDriver("PostgreSQL")
con<-dbConnect(drv,host="localhost",port=5432,dbname="STUDENT",user="postgres",password="postgres")
rs<-dbSendQuery(con,"select * from academic")
dataframe<-data.frame(fetch(rs,n=-1))
dataframe<-data.frame(lapply(dataframe,function(dataframe){replace(dataframe,dataframe==-1,NA)}))
library(e1071)
marks<-read.table('G:\\Academics\\Academics\\academics\\main project\\R_workout\\academic_train.csv' ,header=T,sep=',')
classifier<-naiveBayes(marks[,2:6],marks[,7])
dataframe<-cleandata(dataframe)
df<-data.frame(table(predict(classifier,dataframe,type="class")))
class<-df$Freq
alabel<-round(class/sum(class)*100,2)
alabel<-paste(alabel,"%",sep="")
pie(class,main="A Pie Chart of academic performance of students",col=terrain.colors(3),label=alabel)
legend(1,0.5,c("Elite Performers","Moderate Performers","Poor Performers"),cex=0.8,fill=terrain.colors(3))
}
I am using RCaller for calling a custom R Script. However, I am not sure, how to retrieve result from the custom R code. What should I use? I can run the script using runOnly method. My code is below:
ReplyDeleteimport java.util.Random;
import rcaller.RCaller;
public class Test3 {
public static void main(String[] args){
new Test3();
}
public Test3(){
try{
Random random = new Random();
RCaller caller = new RCaller();
caller.cleanRCode();
caller.setRscriptExecutable("C:\\Software\\R\\R-2.15.3\\bin\\RScript");
caller.R_source("C:\\_Projects\\Integration\\R\\RCodeFrom_Xue\\ForRJava.R");
caller.runOnly();
}catch(Exception e)
{
e.printStackTrace();
//System.out.println(e.toString());
}
}
}
hello
ReplyDeletei have a few problem using Rcalller, iam trying to show a boxplot using Rcaller, but icannt find the syntax
could u guys help me :)
caller.setRscriptExecutable("C:\\Software\\R\\R-2.15.3\\bin\\RScript"); can we give string variable instead of path as argument to setRscriptExecutable method? Does rcaller works well in java serlvet ???
ReplyDeleteHello,
ReplyDeletei have a problem. Anyone use the HilbertVisGUI ever in java calling r? If i runing the program it was flashing up for a second and after that it was close. Can anyone help me? Thanks
code:
c.addDoubleArray("x", numbers);
c.addRCode("library( HilbertVisGUI )");
c.addRCode("hilbertDisplay( x )");
caller.setRCode(c);
caller.runOnly();
Hello
ReplyDeleteWhen i read out the R code by caller.getRCode() method it was return with this:
x<-c(1.0, 200.0, 123.0, 2341.0, 122312.0, 23423.0);
library( HilbertVisGUI )
hilbertDisplay( x )
q("yes")
what's this q("yes") ? and if it was quit how can i turn this into "no"?
please find the new document entry in http://stdioe.blogspot.com/2013/08/a-user-document-for-rcaller.html
ReplyDeletegood luck!
This comment has been removed by the author.
ReplyDeleteI am trying to do same forecasting with java and R and I was wandering is there a way to get the automatically estimated parameters of the forecast model (like HoltWinters (alpha ,beta, gamma)).
ReplyDeleteI try to setup a process of readin a file, then wright out to a new file. But I could not get the ourfile.I don't know why. The following was the code:
ReplyDeleteimport java.io.IOException;
import rcaller.RCaller;
import rcaller.RCode;
public class rJAVA4 {
public static void main(String[] args) throws IOException{
try {
RCaller caller = new RCaller();
RCode code = new RCode();
caller.setRscriptExecutable("C:/Program Files/R/R-2.15.3/bin/Rscript.exe");
code.addRCode("library(Runiversal)");
code.addRCode("x <- read.csv('C:/Documents and Settings/user/桌面/Java sample/example.csv',sep=',',header=TRUE)");
code.addRCode("write.table(x, paste('C:/Documents and Settings/user/桌面/Java sample/grid2.csv',seq=''), quote = FALSE, sep = ',', row.names = T,col.names = T)");
caller.setRCode(code);
}catch (Exception e)
{
System.out.println(e.toString());
}
}
}
the latest version 2.2 of Rcaller does not requires problematic Runiversal package, please visit the page http://stdioe.blogspot.com.tr/2014/04/rcaller-220-is-released.html for details.
ReplyDeletethanks for the rcaller library - very intuitive!
ReplyDeletehowever, am I right assuming that an RCaller instance needs to establish a new connection to R for every e.g. runOnly() command?
As a consequence, multiple RCode objects may not use a single connection but rather:
- either have to be processed with a single e.g. runOnly() command, or
- a new connection between Java and R and thus a new R-session will be established for every single runOnly() command.
I stumble across this issue when trying to execute something like:
RCaller engine = new RCaller();
engine.setRScriptExecutable([path to Rscript.exe]);
// load required R-library
RCode code = new RCode();
code.addRCode("library([lib1])");
engine.setRCode(code);
engine.runOnly();
// load required data file
code.clear()
code.addRCode("source([file1])");
engine.setRCode(code);
engine.runOnly();
// perform calculations
code.clear();
code.addRCode("mean(x)");
engine.setRCode(code);
engine.runAndReturnResults();
This obviously has performance implications in a project where frequent interaction between Java and R is required.
Thanks for your comments!
nils
please have a look at the paper for sequential running of commands within a single R instance:
DeleteSatman, M.H., RCaller: A Software Library for Calling R from Java, British Journal of Mathematics and Computer Science, ISSN: 2231-0851,Vol.: 4, Issue.: 15 (01-15 August), 2014
http://www.sciencedomain.org/abstract.php?iid=550&id=6&aid=4838#.U5XU3PmSy1Y
thanks for the hint. exactly what I was looking for!
Deletenils
can anyone help me why this piece code is not working in RCaller :
ReplyDeletecode : fit <- tslm(datats~trend)
but this working fine in R studio. Please help.
RCaller 2.5 is available in site http://stdioe.blogspot.com.tr/2015/03/rcaller-25-is-available-for-downloading.html
ReplyDeleteHello!!!
ReplyDeletecan you provide the DETAILED descriptions of how to install RCaller in eclipse in windows XP.
step by step installation procedure....... (It would be Very helpful to beginners like me)
because I am looking for it and not getting what i need to know...
Thank you.........
hi ..
ReplyDeleteI am trying to execute below code
package pkg;
import java.io.File;
import javax.swing.ImageIcon;
import rcaller.RCaller.*;
public class Imgcls {
public Imgcls() {
try {
RCaller caller = new RCaller();
// RCode code = new RCode();
caller.setRscriptExecutable("C:\\Program Files\\R\\R-3.2.2\\bin\\i386\\Rscript.exe");
caller.cleanRCode();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
caller.addDoubleArray("x", numbers);
File file = caller.startPlot();
caller.addRCode("plot.ts(x)");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
}
catch (Exception e) {
System.out.println(e.toString());
}
}
public static void main(String[] args) {
new Imgcls();
}
}
Its showing methos addDoubleArray () is undefined in Rcaller
what shall I do now
hi sir
ReplyDeleteI am trying to execute below code It has shown me error
package pkg;
import java.io.File;
import javax.swing.ImageIcon;
import rcaller.RCaller;
public class Imgcls {
public Imgcls() {
try {
RCaller caller = new RCaller();
// RCode code = new RCode();
caller.setRscriptExecutable("C:\\Program Files\\R\\R-3.2.2\\bin\\i386\\Rscript.exe");
caller.cleanRCode();
double[] numbers = new double[]{1, 4, 3, 5, 6, 10};
caller.addDoubleArray("x", numbers);
File file = caller.startPlot();
caller.addRCode("plot.ts(x)");
caller.endPlot();
caller.runOnly();
ImageIcon ii = caller.getPlot(file);
caller.showPlot(file);
}
catch (Exception e) {
System.out.println(e.toString());
}
}
public static void main(String[] args) {
new Imgcls();
}
}
method addDoubleArray(),caller.startPlot(),addRcode(),endPlot() lines showing error as undefined type in Rcaller why?
Can you suggest basics from examples of Rcaller